Phylogenetic turnover quantifies the evolutionary distance among species assemblages and is central to understanding the main drivers shaping biodiversity. It is affected both by geographic and environmental distance between sites.
Category Archives: Senza categoria
Global drivers of population density in terrestrial vertebrates
Although the effects of life history traits on population density have been investigated widely, how spatial environmental variation influences population density for a large range of organisms and at a broad spatial scale is poorly known. Filling this knowledge gap is crucial for global species management and conservation planning and to understand the potential impactContinue reading “Global drivers of population density in terrestrial vertebrates”
A multidisciplinary approach to identify multispecies hotspots of intra-specific diversity
Alice Pezzarossa will be in Jyväskylä (Finland). Read more at https://peerageofscience.org/conference/eccb2018/107814/
Hierarchical, multi-grain rendezvous site selection by wolves in southern Italy
Fine‐scale knowledge of how anthropogenic effects may alter habitat selection by wolves (Canis lupus) is important to inform conservation management, especially where wolf populations are expanding into more populated areas or where human activity and development are increasingly encroaching on formerly pristine environments.
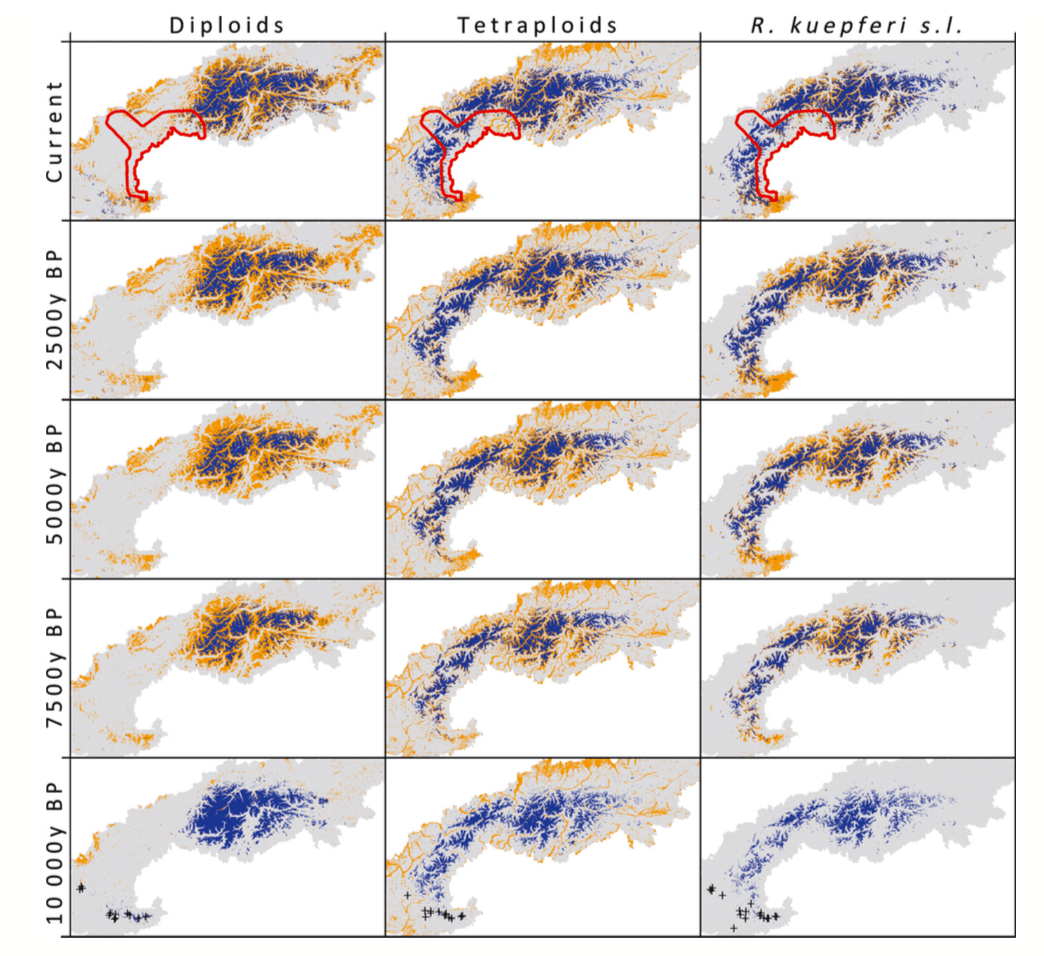
Reconstructing geographical parthenogenesis: effects of niche differentiation and reproductive mode on Holocene range expansion of an alpine plant
Asexual taxa often have larger ranges than their sexual progenitors, particularly in areas affected by Pleistocene glaciations.
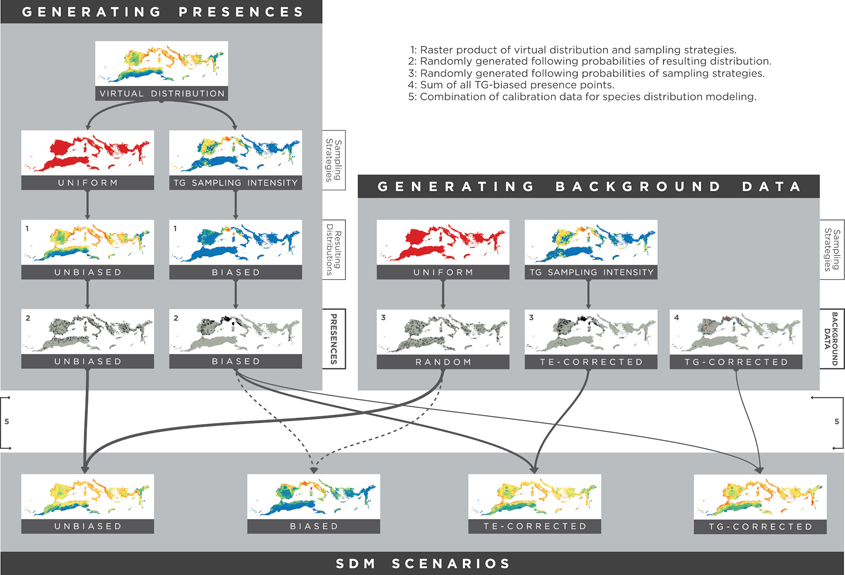
Bias correction in SDMs
Questo è l’estratto per il tuo primo articolo.